You are currently browsing the monthly archive for November 2012.
The Phage strikes back!
For our next discussion I have chosen a paper that will excite, I hope, all of us whether we wave our pom poms for bacteria or viruses alike. Next weeks MicroTWJC will be on: http://dx.plos.org/10.1371/journal.pgen.1003023.t001
I first came across this study when George Salmond recently came to Newcastle to give our Institute seminar and he closed his talk on this story. Then @NatureRevMicro highlighted the paper in a mid-week micro list.
I have followed George’s contributions to microbiology for going on 18-20 years. I first met him in 1997 (or was it 1998 – eek!) when he gave a lecture on an EMBO Advance Eubacterial Genetics course in Umea, Sweden. I remember a quote from this course that I always am reminded of his lecture “have a [transducing] phage, do genetics” as at that time his work centered on Erwinia and was helped by a transducing phage they had isolated.
Other reasons why I have picked this paper include the point that the phage they isolate is in fact a flagellar-specific phage, so it relates to my own field of research. Then we can add my newfound interest in CRISPR systems. As part of our Microbiology degree we recently agreed that we did not do the new interest in phage technology merit. So over the last academic year one new series of lectures I had to prepare was on phages – where I must admit I got quiet enthusiastic about the CRISPR systems. So all in all, I have chosen this paper for many reasons.
Points we should focus on: the general aspects regarding the interaction between bacterial host and phage infection-defence-counter attack. What else would you do in this study? Is there any specifics that stand out for you?
As a bonus (it may not work) we will see if any of George’s lab are on twitter so we may even get author feedback!
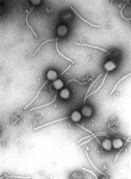
Image:
Photograph courtesy of Vincent Fischetti and Raymond Schuch, The Rockefeller University and released under a Creative Commons Licence here
Next time (Tuesday 20th November) we will be discussing the following paper
Title: Rumen microbial and fermentation characteristics are affected differently by bacterial probiotic supplementation during induced lactic and subacute acidosis in sheep
available from the following link http://www.biomedcentral.com/1471-2180/12/142
Abstract:
Background:
Ruminal disbiosis induced by feeding is the cause of ruminal acidosis, a digestive disorder prevalent in high-producing ruminants. Because probiotic microorganisms can modulate the gastrointestinal microbiota,
propionibacteria- and lactobacilli-based probiotics were tested for their effectiveness in preventing different forms of acidosis.
Results:
Lactic acidosis, butyric and propionic subacute ruminal acidosis (SARA) were induced by feed chalenges in three groups of four wethers intraruminally dosed with wheat, corn or beet pulp. In each group, wethers were either not supplemented (C) or supplemented with Propionibacterium P63 alone (P) or combined with L. plantarum (Lp + P) or L. rhamnosus (Lr + P). Compared with C, all the probiotics stimulated lactobacilli proliferation, which reached up to 25% of total bacteria during wheat-induced lactic acidosis. This induced a large increase in lactate concentration, which decreased ruminal pH. During the corn-induced butyric SARA, Lp + P decreased Prevotella spp. proportion with a concomitant decrease in microbial amylase activity and total volatile fatty acids concentration,and an increase in xylanase activity and pH. Relative to the beet pulp-induced propionic SARA, P and Lr + P improved ruminal pH without affecting the microbial or fermentation characteristics. Regardless of acidosis type, denaturing gradient gel electrophoresis revealed that probiotic supplementations modified the bacterial community structure.
Conclusion:
This work showed that the effectiveness of the bacterial probiotics tested depended on the acidosis type. Although these probiotics were ineffective in lactic acidosis because of a deeply disturbed rumen microbiota, some of the probiotics tested may be useful to minimize the occurrence of butyric and propionic SARA in sheep. However, their modes of action need to be further investigated.
Discussion points:
1. Is the paper well written and easy to follow?
2. Do the authors address the aims of the paper?
3. Was the data conclusive enough?
4. What other experiments could be done?
After a slight hiccup, here is the paper for week 14(ish) of microtwjc! We all have favourite bugs and this one is fast becoming mine (if its cool enough for history guy Dan Snow and C4 filthy cities episode, its good enough for me http://youtu.be/PZgHXAek0No )
Title: Poly-N-Acetylglucosamine Expression by Wild-Type Yersinia pestis Is Maximal at Mammalian, Not Flea, Temperatures
Abstract: Numerous bacteria, including Yersinia pestis, express the poly-N-acetylglucosamine (PNAG) surface carbohydrate, a major component of biofilms often associated with a specific appearance of colonies on Congo red agar. Biofilm formation and PNAG synthesis by Y. pestis have been reported to be maximal at 21 to 28°C or “flea temperatures,” facilitating the regurgitation of Y. pestis into a mammalian host during feeding, but production is diminished at 37°C and thus presumed to be decreased during mammalian infection. Most studies of PNAG expression and biofilm formation by Y. pestis have used a low-virulence derivative of strain KIM, designated KIM6+, that lacks the pCD1 virulence plasmid, and an isogenic mutant without the pigmentation locus, which contains the hemin storage genes that encode PNAG biosynthetic proteins. Using confocal microscopy, fluorescence-activated cell sorter analysis and growth on Congo red agar, we confirmed prior findings regarding PNAG production with the KIM6+ strain. However, we found that fully virulent wild-type (WT) strains KIM and CO92 had maximal PNAG expression at 37°C, with lower PNAG production at 28°C both in broth medium and on Congo red agar plates. Notably, the typical dark colony morphology appearing on Congo red agar was maintained at 28°C, indicating that this phenotype is not associated with PNAG expression in WT Y. pestis. Extracts of WT sylvatic Y. pestis strains from the Russian Federation confirmed the maximal expression of PNAG at 37°C. PNAG production by WT Y. pestis is maximal at mammalian and not insect vector temperatures, suggesting that this factor may have a role during mammalian infection.
Importance: Yersinia pestis transitions from low-temperature residence and replication in insect vectors to higher-temperature replication in mammalian hosts. Prior findings based primarily on an avirulent derivative of WT (wild-type) KIM, named KIM6+, showed that biofilm formation associated with synthesis of poly-N-acetylglucosamine (PNAG) is maximal at 21 to 28°C and decreased at 37°C. Biofilm formation was purported to facilitate the transmission of Y. pestis from fleas to mammals while having little importance in mammalian infection. Here we found that for WT strains KIM and CO92, maximal PNAG production occurs at 37°C, indicating that temperature regulation of PNAG production in WT Y. pestis is not mimicked by strain KIM6+. Additionally, we found that Congo red binding does not always correlate with PNAG production, despite its widespread use as an indicator of biofilm production. Taken together, the findings show that a role for PNAG in WT Y. pestis infection should not be disregarded and warrants further study.
This weeks paper can be found here http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3419525/pdf/mBio.00217-12.pdf
Usual questions apply…..
- Is this paper well written?
- Does the data support the conclusion?
- What other data/experiments would you like to have seen presented? and most importantly
- Where next to determine a role (if there is one?!?) for PNAG in Y. pestis virulence?
Hope everyone can make it to #microtwjc on Tuesday